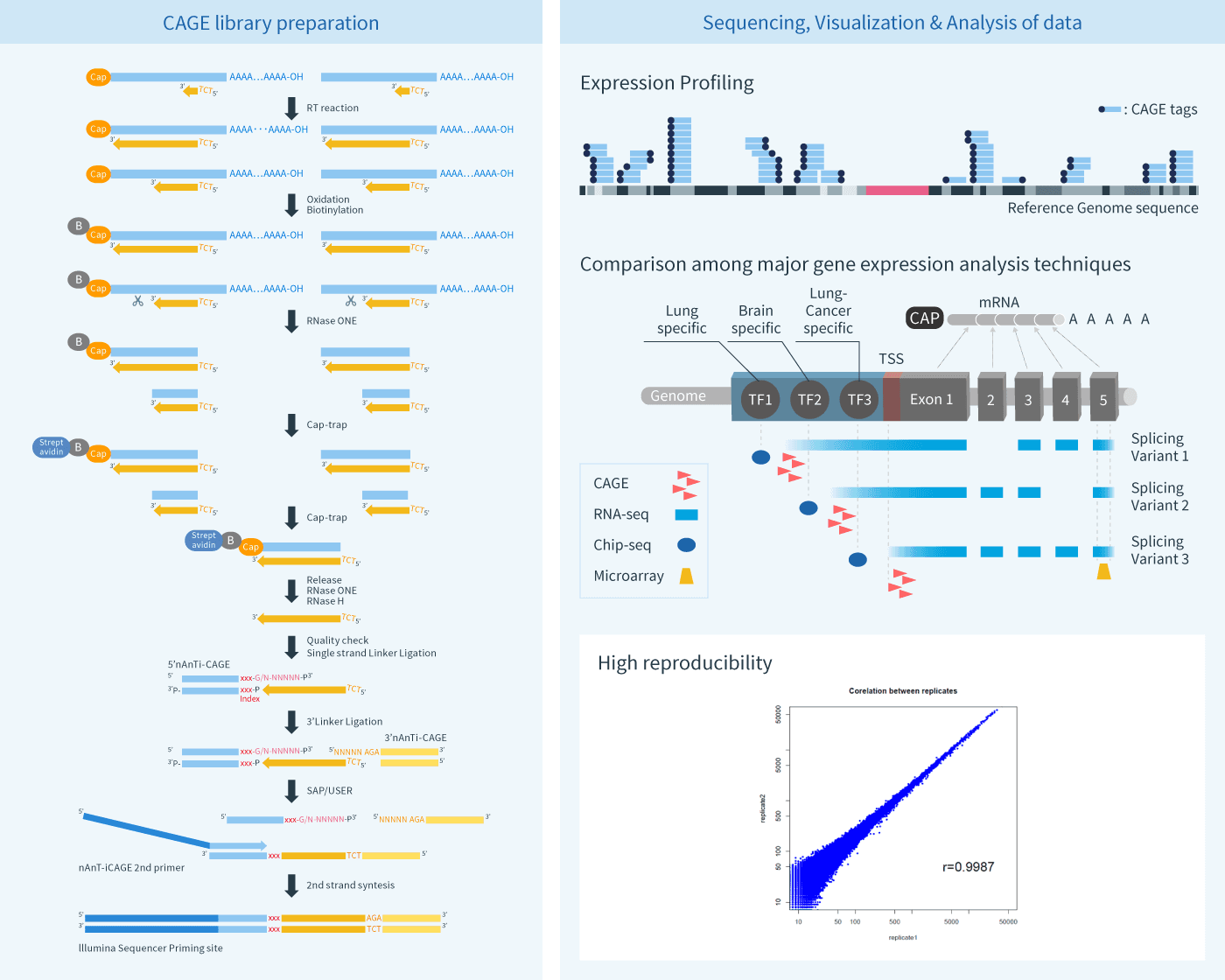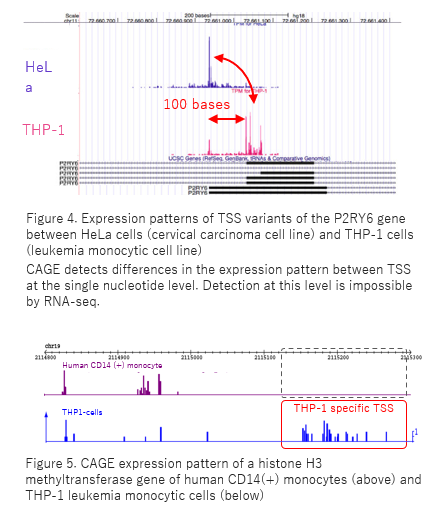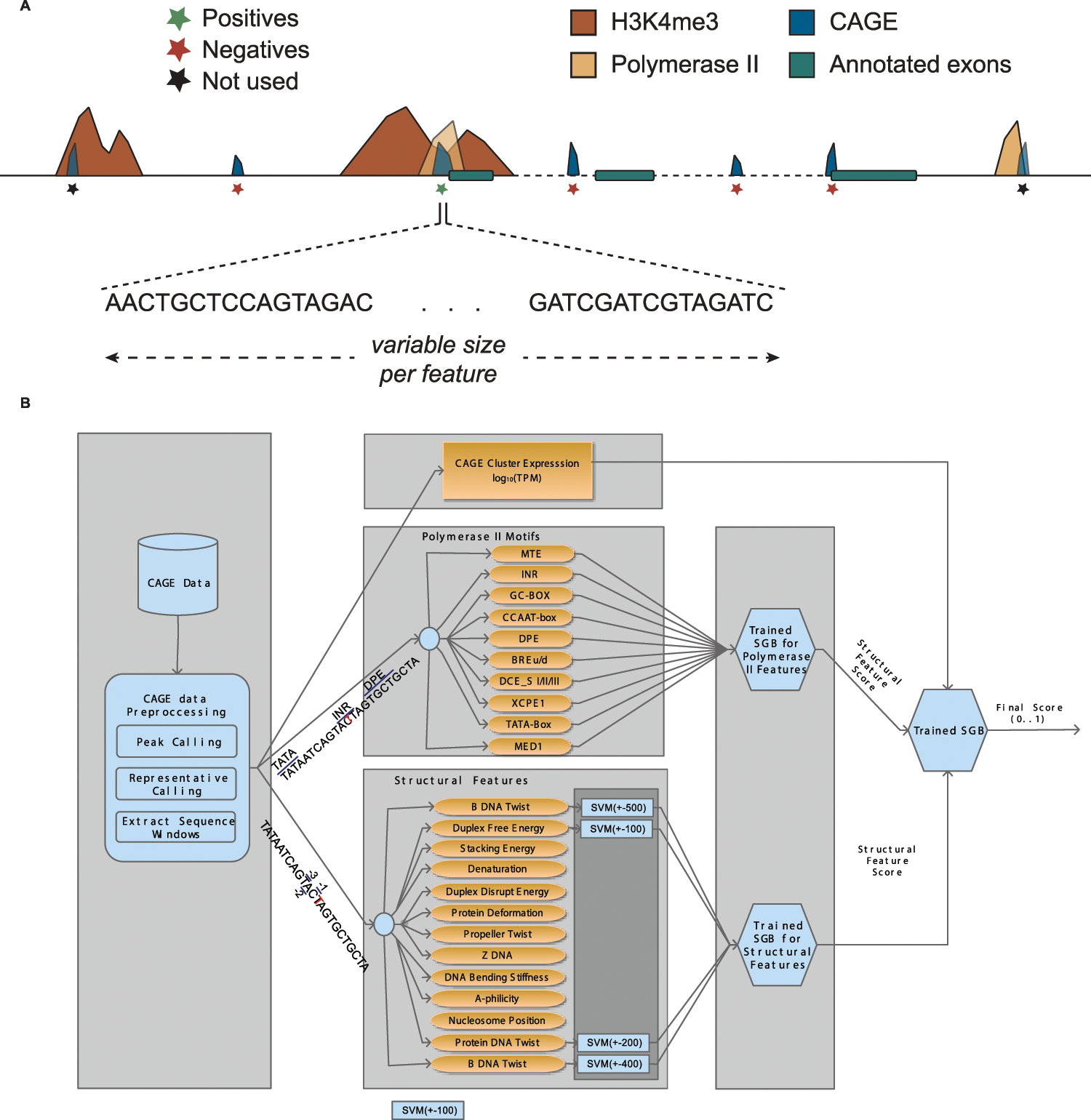
Solving the transcription start site identification problem with ADAPT-CAGE: a Machine Learning algorithm for the analysis of CAGE data | Scientific Reports

Nascent RNA sequencing analysis provides insights into enhancer-mediated gene regulation | BMC Genomics | Full Text
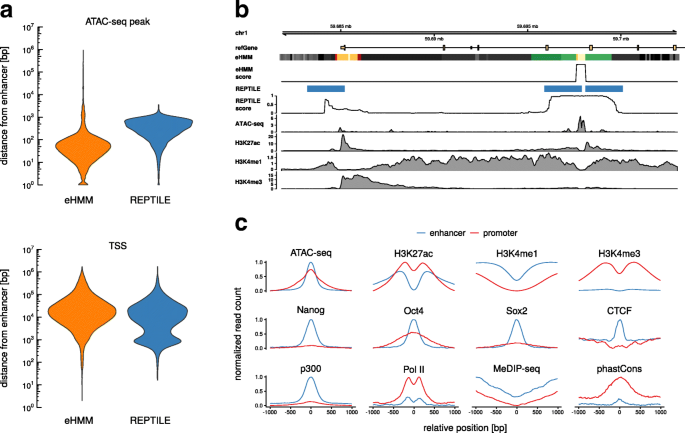
Predicting enhancers in mammalian genomes using supervised hidden Markov models | BMC Bioinformatics | Full Text
ChIP-GSM: Inferring active transcription factor modules to predict functional regulatory elements | PLOS Computational Biology

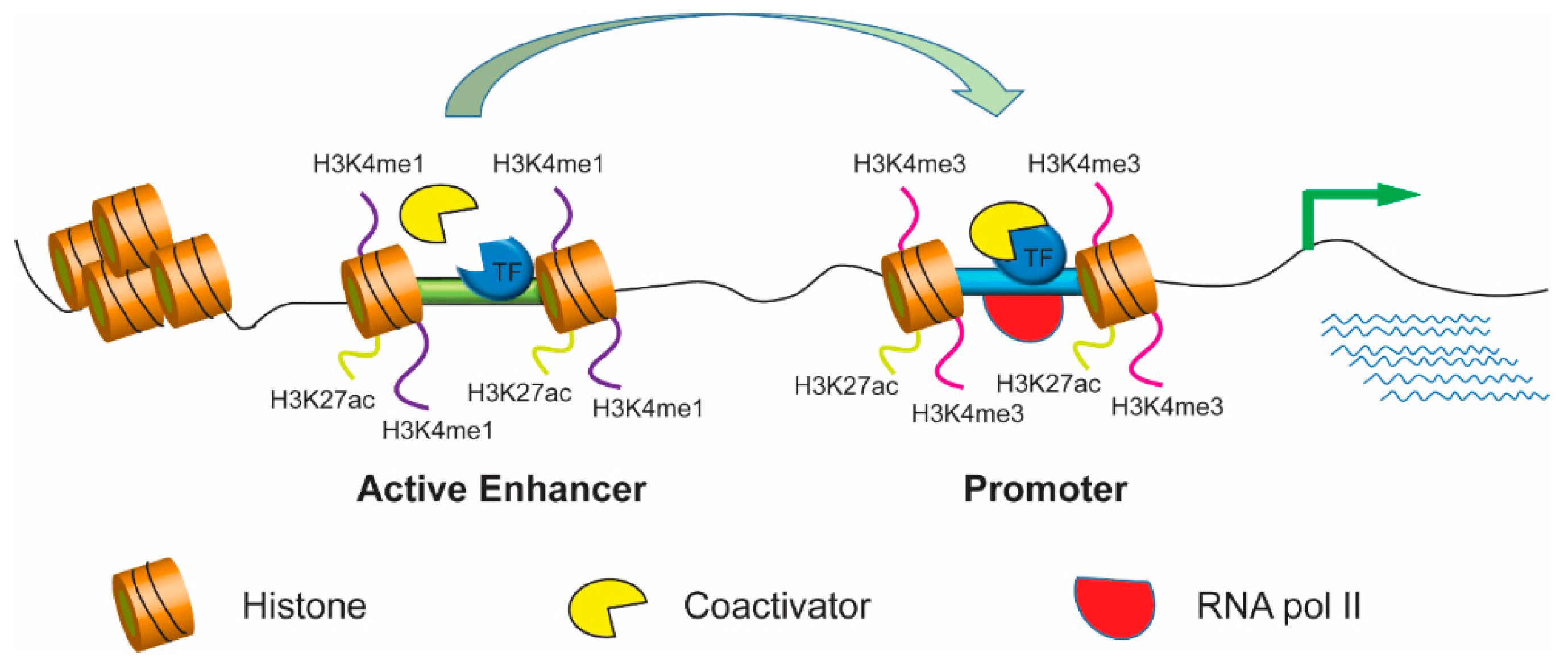
Enhancer and super‐enhancer: Positive regulators in gene transcription - Peng - 2018 - Animal Models and Experimental Medicine - Wiley Online Library
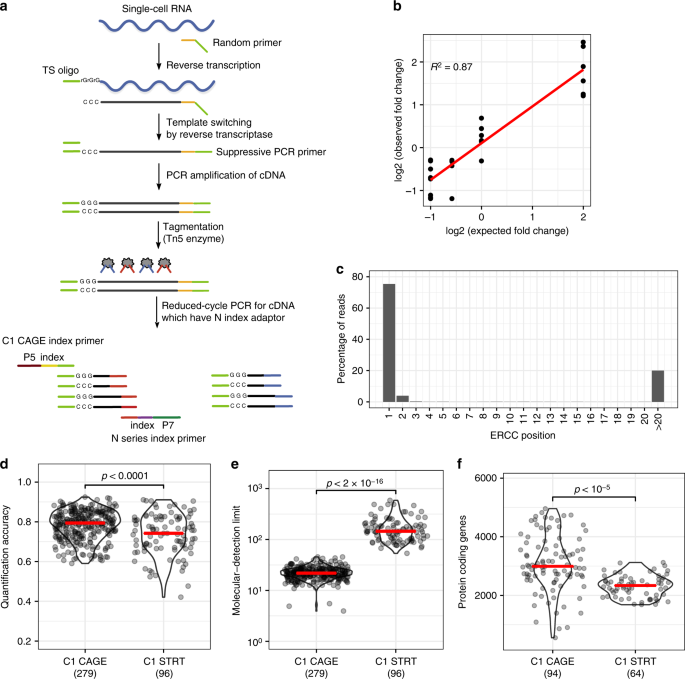
C1 CAGE detects transcription start sites and enhancer activity at single-cell resolution | Nature Communications
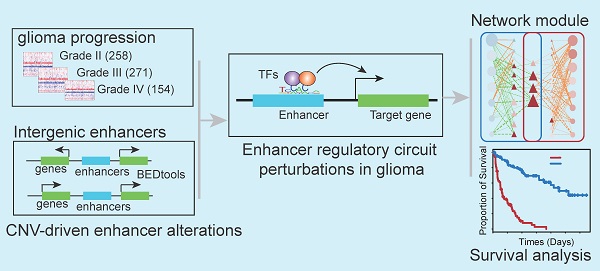
Systematic analysis of enhancer regulatory circuit perturbation driven by copy number variations in malignant glioma

A High-Resolution Map of Human Enhancer RNA Loci Characterizes Super- enhancer Activities in Cancer - ScienceDirect

Frontiers | Mapping Mammalian Cell-type-specific Transcriptional Regulatory Networks Using KD-CAGE and ChIP-seq Data in the TC-YIK Cell Line

De novo identification of enhancers with higher sensitivity in NET-CAGE... | Download Scientific Diagram
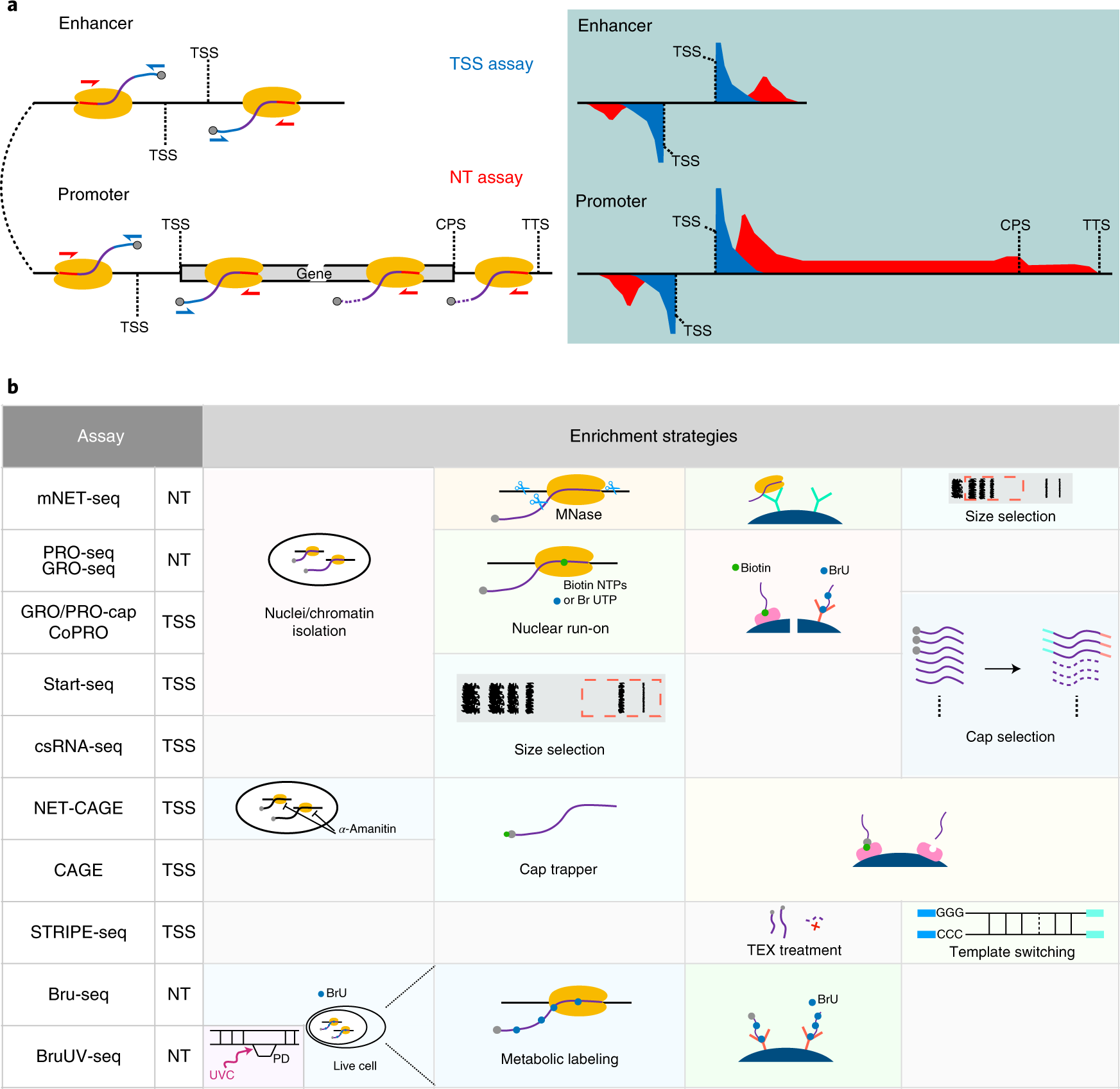
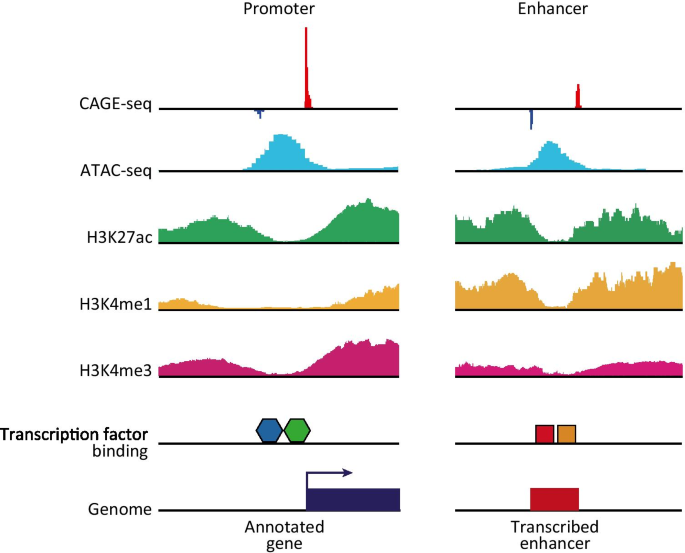
A comparison of experimental assays and analytical methods for genome-wide identification of active enhancers | Nature Biotechnology